picture:
To address this challenge, the team identified low-intratumor heterogeneity/high-intertumor variability (LIHV) genes. This signature of 1,341 genes targets markers that remain consistent within a human tumor but vary greatly between patients. By filtering out the discrepant “noise”, the LIHV design enables a robust and reproducible classification of iCCA into five molecular subtypes, each associated with a unique therapeutic disadvantage:
vision Again
Credit: BGI Genomics & Zhongshan Hospital of Fudan University
Intrahepatic cholangiocarcinoma (iCCA), the second most common liver cancer after hepatocellular carcinoma, remains one of the most difficult diseases to treat due to its highly hidden and heterogeneous nature. A collaborative team from BGI Genomics and Fudan University’s Zhongshan Hospital has now mapped the molecular “chaos” of iCCA, providing a roadmap for more accurate diagnosis and treatment. The research is printed on Cell Reports Medicine.
A 1,341-gene signature to reduce sampling bias
A major obstacle in the treatment of iCCA is the prevalence of intratumoral heterogeneity. Cancer cells from different areas of the same tumor can show unique molecular characteristics, meaning that a single biopsy often fails to capture the full biological environment. Researchers have found that the latest classification methods identify more than a quarter of tumors and two-thirds of the reports of the immune system, depending on the location of the biopsy.
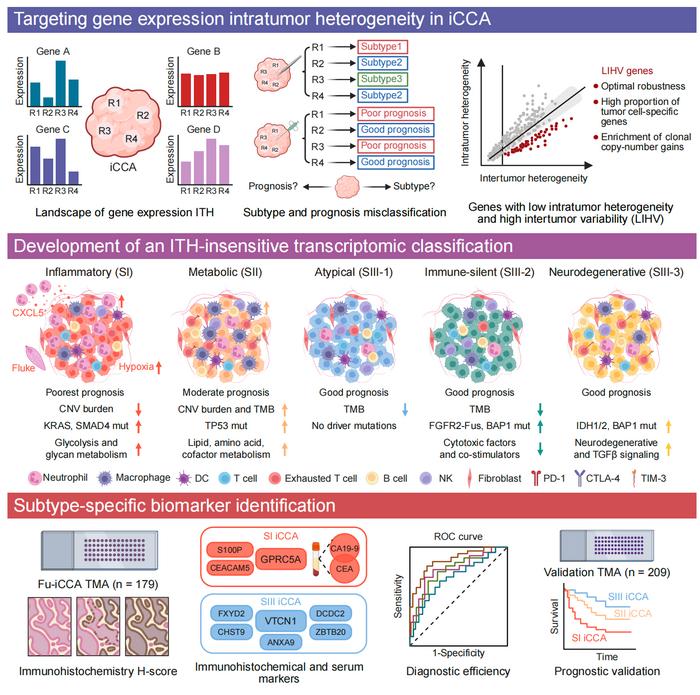
To address this challenge, the team identified low-intratumor heterogeneity/high-intertumor variability (LIHV) genes. This signature of 1,341 genes targets markers that remain consistent within a human tumor but vary greatly between patients. By filtering out the discrepant “noise”, the LIHV design enables a robust and reproducible classification of iCCA into five molecular subtypes, each associated with a unique therapeutic disadvantage:
Subtype of inflammation (SI): It is associated with the poorest prognosis and is enriched in the large bile duct-type iCCA. These tumors are usually permanent KRAS and SMAD4 changes and show elevated levels of serum CA19-9 and CEA. Their microenvironment is characterized by a strong neutrophil infiltration, and the chemokine CXCL5 plays a central role.
Sub-metabolic type (SII): It shows the highest burden of tumor transformation and the highest expression of many molecular diagnostics. Despite the immunosuppressive phenotype, this profile suggests potential for immune checkpoint inhibitors and combination immunotherapies.
Group SIII (SIII-1, SIII-2, SIII-3): Consisting of atypical, immune-selent, and neurodegenerative subtypes, respectively, this group is most associated with the small bile duct-type iCCA and generally shows better clinical outcomes. Various methods of genetic modification are observed, including enhancement BAP1 changes in SIII-2 and IDH1/2 changes in SIII-3.
Although previous studies have classified iCCA based on genetic changes or aspects of the tumor microenvironment, this design specifically emphasizes genes that remain stable in tumors, thus reducing sampling bias and improving classification reliability.
Converting subtypes to ttreatment smethods
Regardless of classification, the real clinical value of this framework lies in its ability to guide treatment decisions. For subtype of inflammation (SI).highly aggressive and often resistant to PD-1 inhibition, the study shows for the first time that HSP90 inhibitors can suppress tumor growth. Importantly, these agents appear to sensitize tumors to PD-1 therapy, suggesting a promising combination strategy for high-risk patients. To neurodegenerative (SIII-3) subtypetumors show high expression of the immunological test TIM-3. Experimental evidence shows that dual inhibition of PD-1 and TIM-3 is more effective in controlling tumor proliferation than single therapy. The metabolic (SII) subtype presents a paradox: despite being an immunosuppressive phenotype, it produces high levels of many diagnostic molecules. This suggests that combination therapies targeting CTLA-4 and LAG3 may provide an effective strategy to overcome immune resistance. Although these studies are currently in their infancy, they form the basis for future clinical trials tailored to specific molecular profiles.
Functional Biomarkers for Clinical Application
To aid clinical translation, researchers have identified available biomarkers that can serve as alternative markers for each type.
GPRC5A was identified as a specific marker for the inflammatory subtype (SI), showing a sensitivity of 71.4% and a specificity of 88.9%. This marker can help pathologists identify this aggressive form in routine histopathological examination.
VTCN1 (also known as B7-H4) was identified as a marker for group SIII, with a sensitivity of 68.7% and a specificity of 80.6%. In addition, the serum levels of CEA and CA19-9 can serve as non-invasive indicators of the underlying type of inflammation, which supports the integration of molecular features in the flow of clinical activity.
Towards Precision Oncology at iCCA
This study provides a comprehensive “map” to navigate the complexity of iCCA by directly linking molecular subtypes to prognosis and targeted therapies. The LIHV gene signature effectively reduces the limitations imposed by tumor heterogeneity, enabling small biopsy samples to accurately reflect the entire tumor. Future work will focus on validating this design in different patient populations and developing treatments guided by a clinical trial model. By moving beyond single methods to accurate data oncology, this map holds the promise of significantly improving survival outcomes and quality of life for patients with iCCA.
About BGI Genomics
BGI Genomicsheadquartered in Shenzhen, China, is the world’s leading provider of integrated precision medicine solutions. Our services cover more than 100 countries and territories, including more than 2,300 medical centers. In July 2017, as a subsidiary of BGI Group, BGI Genomics (300676.SZ) was officially listed on the Shenzhen Stock Exchange.
Journal
Cell Reports Medicine
Publication Date of Articles
30-Mar-2026
Description: AAAS and EurekAlert! are not responsible for the accuracy of the content presented on EurekAlert! by participating in organizations or for the use of any information through the EurekAlert system.
#genetic #mapping #improves #optimal #treatment #intrahepatic #cholangiocarcinoma